The Discovery Engine Program
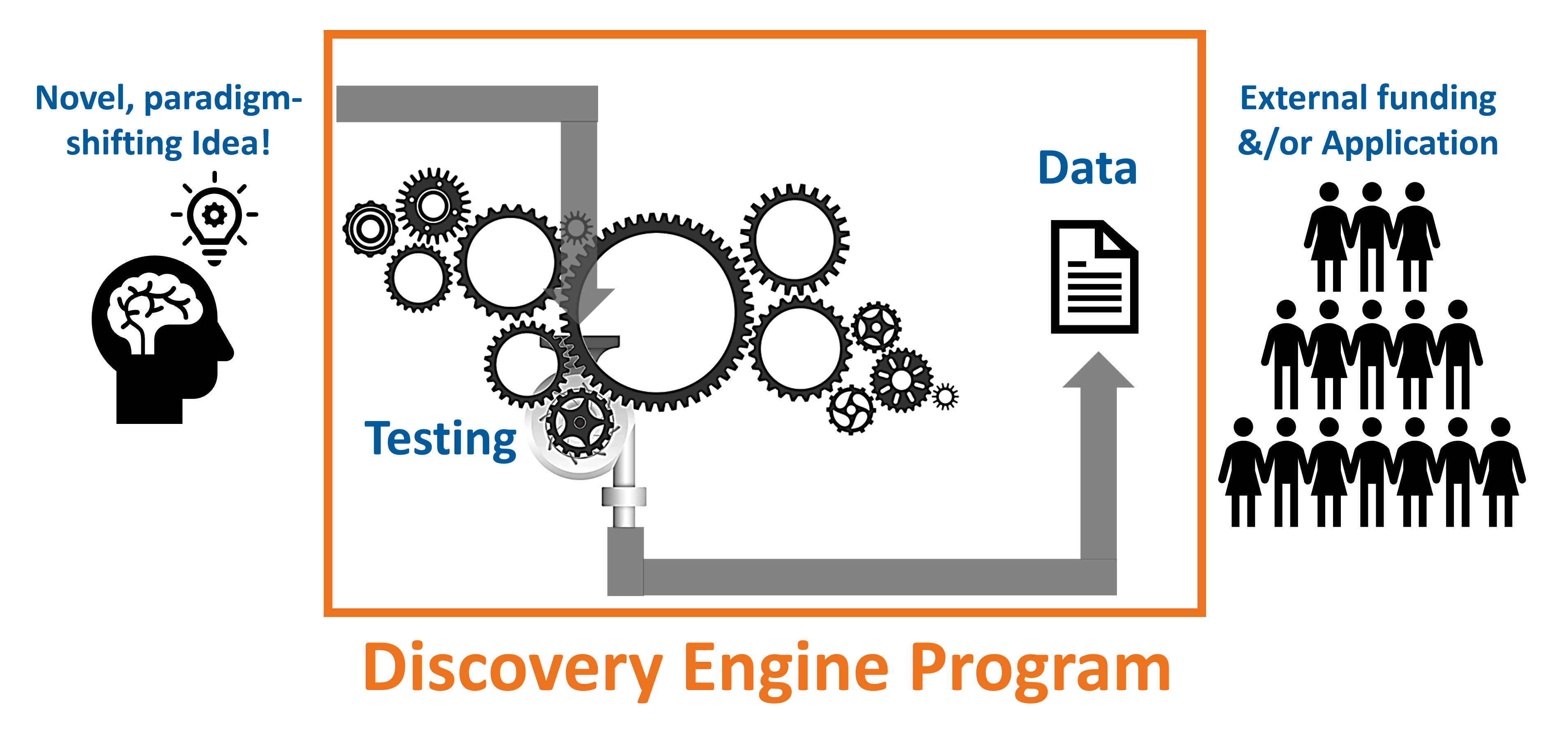
The Discovery Engine Program was designed to foster ideas and provide a sheltered environment to test these ideas with real funding. We seek proposals that go beyond the applicant mentor’s research premise in new directions.
Who can apply?
Any non-faculty member of the Lyda Hill Department of Bioinformatics, non-faculty lab members of PIs with secondary appointments in Bioinformatics, and graduate students of the Computational Biology program.
What should you submit?
- A single page document that addresses each of the four questions below. Formatting: Page margins 0.5 inches on all sides, 11-point Arial font, single spaced.
- What is the idea?
Criteria to bear in mind- Idea should be your own, without influence or contribution from your mentor
- Should be distinct from but may build on current research
- Should not have been submitted to any non-departmental UTSW or external funding opportunities
- What possible discovery will come from it? (End goal)
- Who will care about this discovery? (Audience, impact and significance)
- Why are you the first to have this idea? (Rationale)
How was the idea formed? Why has your mentor not pursued this direction of research? What propelled your thinking in this new direction?
- What is the idea?
- A budget (no page limit)
- Justification of your budget (no page limit)
- Endorsement from your PI and each of your collaborators. Each MUST contain statements of feasibility of proposed project and a testimonial of independence.
Formatting: each endorsement restricted to half page, 0.5 inch margins on all sides, 11-point Arial font, single-spaced. For example, if you have 2 collaborators, you would submit 1.5 pages total containing 3 endorsement statements. If you have no collaborators, you would submit a half page endorsement statement from your PI.
You may email these materials to Xiujuan (Joanna) Han. Please collate documents in above specified order in one PDF.
Next deadline: June 1, 2026 (cutoff 11.59 p.m.)
Applications will be screened by primary Bioinformatics faculty (Tenured/Tenure-track, Research-Track faculty at rank of Assistant Professor and higher, and UTSW Distinguished Fellow). All applications will be reviewed for idea soundness and financial feasibility. If necessary, a pitch session will be arranged. Decision and commitment letters will be sent out within one month after deadlines.
Successful Awards
March 2024 Award
Gang-of-N Decoding – Inference-Time Ensembling of Large Language Models
Submitted by Michael Holcomb, M.S. (Jamieson Lab)
June 2023 Award
A Bright Future: Introducing light-based allosteric control to cytochrome p450
Submitted by James McCormick, Ph.D. (Reynolds Lab)<
March 2023 Award
Developing platelet profiling-based diagnosis tools for Alzheimer's Disease
Submitted by Muhammed Sadik Yildiz, Ph.D. (Toprak Lab)
