UTSW researchers identify first Beta variant in North Texas amid rise of "variants of concern"
DALLAS – June 17, 2021 – UT Southwestern Medical Center scientists have identified the first two cases of the Beta variant (B.1.351) of COVID-19 infection in North Texas using next-generation sequencing technologies along with targeted PCR testing.
Also of note, the most recent UT Southwestern sequencing demonstrates increasing dominance of “variants of concern” – Alpha (B.1.1.7), Beta (B.1.351) and Delta (B.1.617). These variants of concern accounted for 83 percent of variants for the week of May 28 through June 3, compared with 75 percent the prior week and 55 percent the week before that.
“As variants of concern continue to be detected in North Texas and increasingly dominate, it is more important than ever for individuals to strongly consider or reconsider vaccination,” says James Cutrell, M.D., associate professor of internal medicine and program director of UTSW’s Infectious Diseases Fellowship Program. “The encouraging news is that the approved vaccines appear to be holding up against the variants and do help slow the transmission of all types of virus while protecting against more severe disease.”
UT Southwestern sequencing protocols show the Alpha variant (B.1.1.7) remains dominant in North Texas – found in about 60 percent of sampled individuals – followed by a rise in the Delta variant (B.1.617), now in more than 20 percent of sampled individuals. The Beta variant (B.1.351) was found in about 4 percent of individuals sampled. No individuals tested positive this week for the Gamma (P.1) variant found earlier, Epsilon (B.1.429/B.1.427) or the Iota variant (B.1.526), however all but the Epsilon variant have been previously sampled.
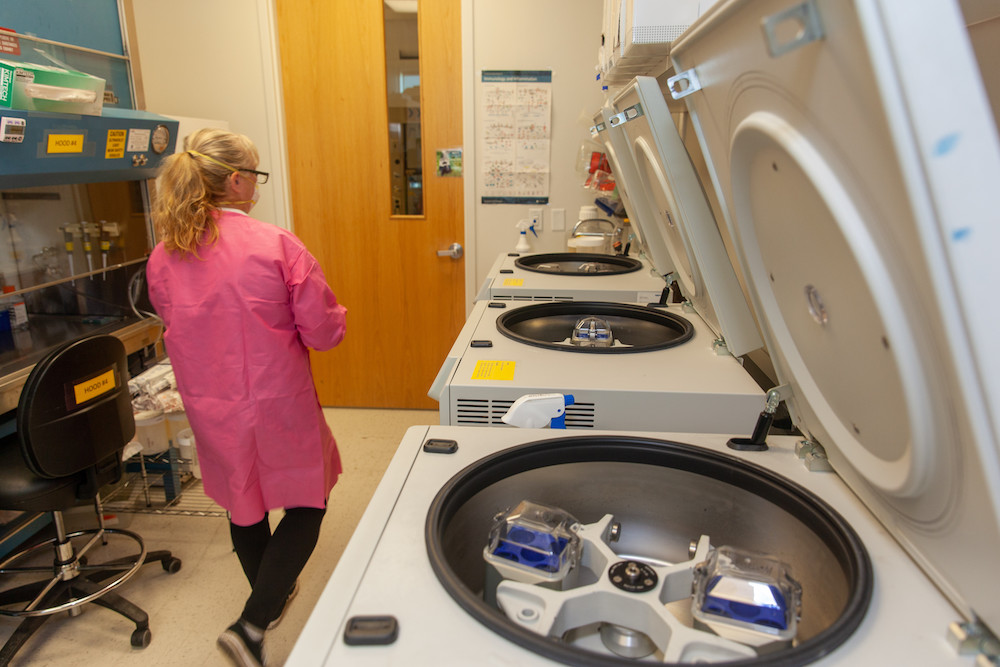

For all COVID-19-positive samples received in UTSW’s clinical lab in the last few months, UT Southwestern has sequenced the virus in order to detect any known variants as well as potential novel new ones – currently about 120 samples per month. Although a relatively small sample, ongoing sequencing gives a better picture of how frequent each of the variants are, and the prevalence of emerging variants such as the Beta variant this week. UT Southwestern reported these findings to Dallas County Health and Human Services (DCHHS).
The growing presence of the Delta variant in North Texas – which more than doubled to 21 percent in sampled individuals the week of May 28 through June 3 – is of concern because it is more transmissible than the original variant and may cause more severe disease. The Delta variant, which accounts for about 10 percent of national cases now, is projected by some models to become the dominant variant in the U.S.
Because each variant has different rates of transmissibility, identifying the different variants appearing in North Texas is important for accurate modeling to predict prevalence, which in turn helps health care officials better prepare for surges, increased hospitalizations, and needed resources. Prevalence of variants can also influence treatment.
| WHO Designation | Pango Lineage | First Identified | Variant Category |
|---|---|---|---|
| Alpha | B.1.1.7 | United Kingdom (09-2020) | Concern |
| Beta | B.1.351 | South Africa (05-2020) | Concern |
| Gamma | P.1 | Brazil (11-2020) | Concern |
| Delta | B.1.617.2 | India (10-2020) | Concern |
| Epsilon | B.1.427/B.1.429 | USA (03-2020) | Interest |
| Zeta | P.2 | Brazil (04-2020) | Interest |
| Eta | B.1.525 | Multiple (12-2020) | Interest |
| Theta | P.3 | Philippines (01-2021) | Interest |
| Iota | B.1.526 | USA (11-2020) | Interest |
| Kappa | B.1.617.1 | India (10-2020) | Interest |
The sequencing was performed in the McDermott Center Next Generation Sequencing (NGS) Core with analysis performed by the McDermott Bioinformatics Lab, both under the supervision of Helen H. Hobbs, M.D., a Howard Hughes Medical Institute investigator and professor of internal medicine and molecular genetics, who directs the Eugene McDermott Center for Human Growth and Development at UT Southwestern. Jeffrey SoRelle, M.D., assistant instructor of pathology, is collaborating with Hobbs by providing all positive COVID-19 samples tested at UT Southwestern and corroborating results with a rapid, focused PCR-based test developed in the Once Upon a Time Human Genomics Center. The collaboration with the McDermott Center Next Generation Sequencing (NGS) Core at UT Southwestern, allows whole genome whole sequencing of SARS-CoV-2 virus in a state-of-the-art facility that performs NGS coupled with bioinformatic analysis. Whole genome sequencing is the best way to monitor for any new variants. Variant data will be integrated into forecasting models for epidemiologic predictions.
Hobbs holds the Philip O'Bryan Montgomery, Jr., M.D. Distinguished Chair in Developmental Biology, the Eugene McDermott Distinguished Chair for the Study of Human Growth and Development, and the 1995 Dallas Heart Ball Chair in Cardiology Research.
About UT Southwestern Medical Center
UT Southwestern, one of the nation’s premier academic medical centers, integrates pioneering biomedical research with exceptional clinical care and education. The institution’s faculty members have received six Nobel Prizes and include 24 members of the National Academy of Sciences, 25 members of the National Academy of Medicine, and 13 Howard Hughes Medical Institute Investigators. The full-time faculty of more than 3,300 is responsible for groundbreaking medical advances and is committed to translating science-driven research quickly to new clinical treatments. UT Southwestern physicians in more than 80 specialties care for more than 143,000 hospitalized patients, attend to more than 470,000 emergency room cases, and oversee nearly 5.3 million outpatient visits a year.
