Study reveals how viruses hijack cells to give them an edge in invasion
UTSW finding could open door to development of drugs to block this process
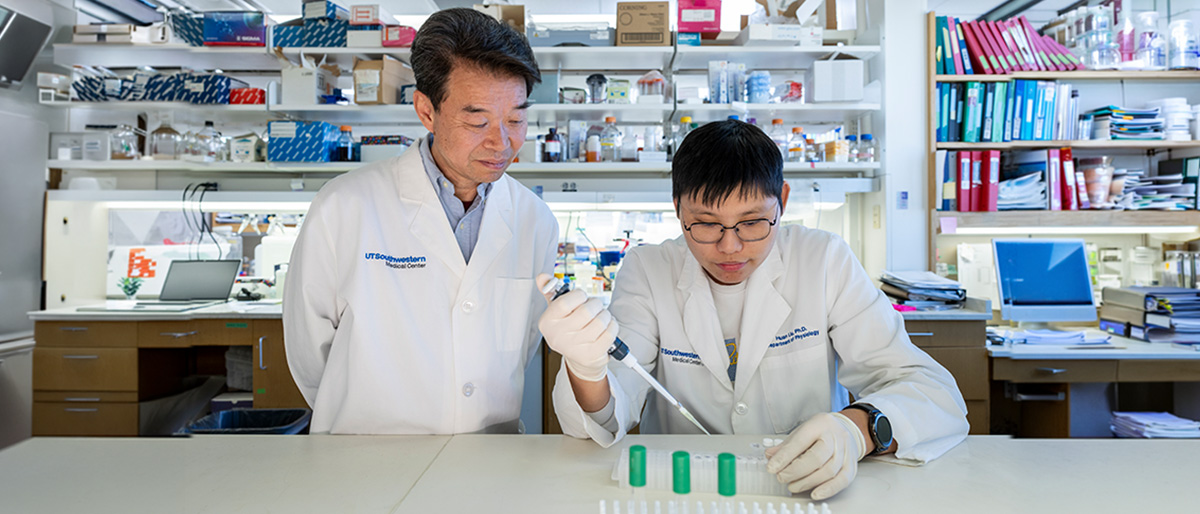

UT Southwestern researchers have found that diverse human viruses have a specialized structure on their RNA that allows them to bypass their host cells’ quality-control constraints. This evasion allows viruses to efficiently produce proteins within hijacked host cells, even though viruses have lower-quality genetic coding.
RNA is a nucleic acid essential for gene expression that carries genetic instructions from DNA to ribosomes for protein synthesis.
“Our finding suggests a new way to block viral protein production,” said Yi Liu, Ph.D., Professor of Physiology and senior author of the study, which appeared in Nature. “If future drugs can force viral RNAs into a looped shape or otherwise interfere with the way viral RNAs are read by the cell, we might selectively slow down virus protein production without shutting down normal cell functions.”

The core of the problem lies in segments of RNA called codons. RNA’s genetic code – represented by the letters A, C, G, U – combines in triplets called codons to specify the amino acids that make up proteins. For instance, the codon GCU “spells” the amino acid alanine. There are 64 possible codons, and only 20 amino acids found in various organisms, so most amino acids can be encoded by two to six codons. For example, alanine can be coded by four “synonymous” codons: GCU, GCC, GCA, and GCG.
However, synonymous codons aren’t used at the same frequencies in all organisms – a phenomenon called codon usage bias. Dr. Liu’s team compared codon usage in 26 families from more than 1,000 viruses that cause disease in humans. From the coronavirus that causes COVID-19 to those causing hepatitis, codon usage of almost all human viruses is skewed toward poorly spelled codons ending in U or A, researchers found. Paradoxically, these viruses can produce proteins just as well as properly spelled host RNA to cause diseases or pandemics.
“The question is, how can they express their proteins so efficiently when they have such poor codon usage?” Dr. Liu said.
The answer was found in different structures of human and viral RNA. At human RNA’s 5′ (five prime) end – the end that feeds into the protein-making machinery – there’s a “cap” that includes a modified G unit.
When human RNA is at work, the 5′ cap interacts with the 3′ end to form a circle. But viruses, instead of a cap, have a variety of loops and hairpin turns that turn their 5′ end into a bulky tangle. And during protein synthesis, the viral RNA remains linear instead of looping.
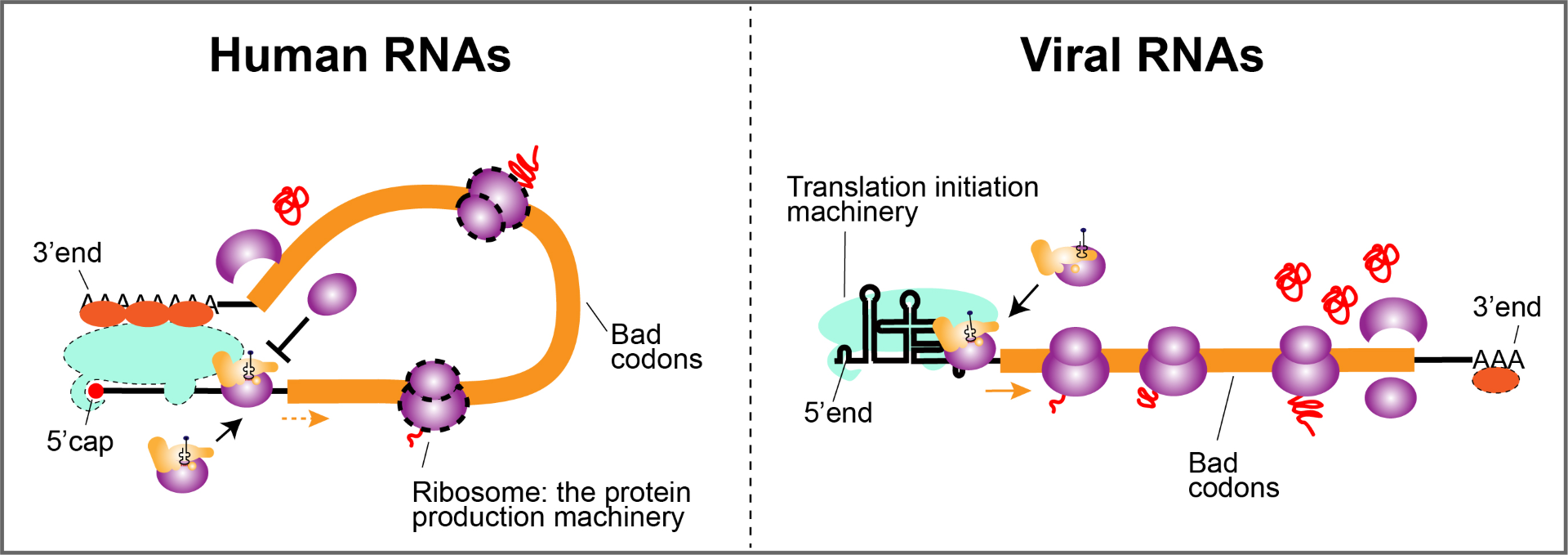
Dr. Liu’s team discovered that straight RNA segments neutralize the host cell’s bias against badly spelled codons so that the viral proteins are produced as efficiently as host proteins.
The researchers then modified the mRNAs with viral 5′ end regions to form circles and found that the host cell could strongly suppress protein production. This showed that circularization is a key step in determining protein synthesis efficiency.
“When viral RNAs were made like normal human mRNAs by forcing them to loop, the bad codons started to matter again and viral protein production dropped dramatically,” Dr. Liu said.
Should further research confirm a way to interfere with this viral RNA activity, Dr. Liu said, a path forward for drug development would be the next phase of research. If successful, this holds potential to combat a variety of viruses, including SARS-CoV-2, the virus that causes COVID-19.
UT Southwestern researchers contributing to this study include Huan Liu, Ph.D., and Jiabin Duan, Ph.D., postdoctoral fellows, Renu Garg, Ph.D., Research Scientist, and Pancheng Xie, Ph.D., Instructor of Physiology. The work was supported by the National Institutes of Health and Lyda Hill Philanthropies.
Endowed Titles:
Dr. Liu holds the Nancy B. and Jake L. Hamon Distinguished Chair in Basic Cancer Research and is a Louise W. Kahn Scholar in Biomedical Research.


